8th Edition of International Conference on Neurology and Neurological Disorders 2023 Report:
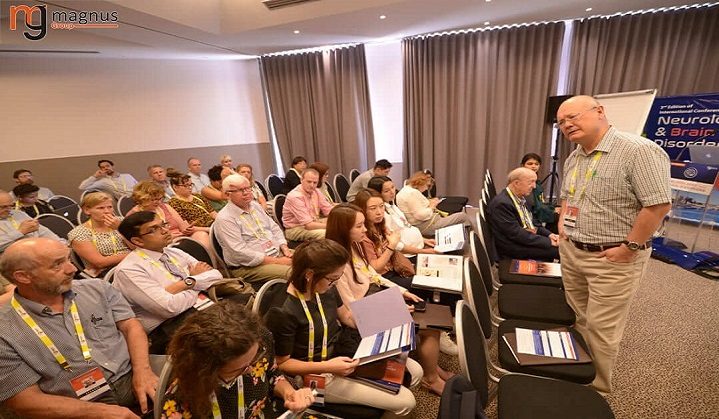
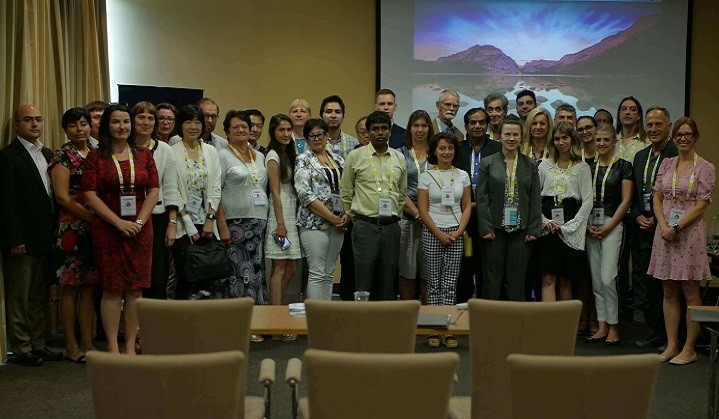
Magnus Group gleefully announces the successful outcome of “8th Edition of International Conference on Neurology and Neurological Disorders” (INBC 2023) which took place on October 19-21, 2023 in Boston, USA.
The congress ran on the theme “BRAIN: Bolstering & Renovating Advanced techniques In Neurology”
INBC 2023 offered an advanced forum for discussing and deliberating the most recent discoveries and clinical approaches in brain healthcare. This encompassed diagnostic techniques, monitoring tools, and management strategies geared towards enhancing neurodevelopmental results. The event delved into methodologies for understanding, safeguarding, and nurturing both developing and mature brains. Over the course of three days, the conference facilitated opportunities for ongoing medical education, catering to a diverse range of professionals dedicated to enhancing the neurological and developmental well-being of patients with neurological conditions
For INBC 2023 Final Program: Click Here
For INBC 2023 Abstract Book: Click Here
For INBC 2023 Gallery: Click Here
We would like to extend our sincere gratitude to the Keynote Speakers, Speakers, Moderators, Poster Presenters, Students, and Delegates for their participation and contribution to the event's success.
Hopefully, you enjoyed both the scientific and networking aspects of the event and benefited from the chance to expand your current profession. I am optimistic that the vast majority of you will continue to work with me in the near future.
Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference | Brain Congress 2024 | International Neuroscience Conference 2024
Organizing Committee INBC 2023:
| Ken Ware |
NeuroPhysics Therapy Institute and Research Centre |
Australia |
| Roberto Di Maio |
University of Pittsburgh |
United States |
| Rocco Gennaro |
University of Southern Indiana |
United States |
| Juan Moreira |
CNC / Gnosis Neurointegrative Center |
Puerto Rico |
| Yong Xiao Wang |
Albany Medical College |
United States |
| Jun Hua |
Johns Hopkins University School of Medicine |
United States |
| Roy G Beran |
School of Medicine at Griffith University |
Australia |
| Ulrich Sprick |
Alexius/Josef Clinic |
Germany |
| Brandon Lucke Wold |
University of Florida |
United States |
Magnus Group is prerogative to thank the Organizing Committee Members, Keynote Speakers; Chair and Co-chairs on engross the plenary sessions, workshops, and special sessions in an expanded manner to make this conference a privileged Summit.
INBC 2023 Speaker Line Up:
Day 1: In-Person-Keynote Speakers:
| Ulrich Sprick |
Alexius/Josef Clinic |
Germany |
| Thomas J. Webster |
Interstellar Therapeutics |
United States |
| Mohammad Zare |
UTHealth and Harris Health System |
United States |
| Jennifer LaHue, Kathryn Crary |
Harris Health System |
United States |
| Ken Ware |
NeuroPhysics Therapy Institute |
Australia |
Day 1: In-Person-Speakers:
| Bharath Kumar |
Nagaraj, Revature LLC |
United States |
| Naveen Kunchakuri |
EchoStar Corporation |
United States |
| Nina Sherman |
Podcaster |
United States |
| Alexis Tang |
TCU School of Medicine |
United States |
| Jag H Khalsa |
GWU School of Medicine and Health Sciences |
United States |
| Arvinder |
AIIMS |
India |
| Hiroko Ikeshima-Kataoka |
Keio University |
Japan |
| Mia W McNary |
Artist and Visual Storyteller, Picture Recovery |
United States |
| Andrew H Hahn |
Life Centered Therapy Training Institute |
United States |
WORKSHOP:
| Ulrich Sprick |
Alexius/Josef Clinic |
Germany |
| Leandro Heidy Yoshioka |
University of Sao Paulo |
Brazil |
| Marta Imamura |
University of Sao Paulo |
Brazil |
| Gilson Tanaka Shinzato |
Clinic of the Medical Faculty, Universtity of Sao Paulo |
Brazil |
| Thi-Lan Freedman |
STORZ Medical |
Switzerland |
Day 1: In-Person-Posters:
| Mikaela Atkins |
Arizona State University |
United States |
| Shehata Anwar |
University of Illinois at Urbana Champaign |
United States |
| Jinyan Zhou |
University of Illinois at Urbana Champaign |
United States |
| Cay Anderson-Hanley |
Union College & iPACES |
United States |
| Jariya Umka Welbat |
Khon Kaen University |
Thailand |
| Apiwat Sirichoat |
Khon Kaen University |
Thailand |
| Edlira Shemsi |
Regional Hospital Durres |
Albania |
| Amy Zhou |
University of Saskatchewan |
Canada |
| Dominique Hayduk Montecino |
Lakeland Regional Health |
United States |
| Alexander Guo |
Timberline High & Boise State University |
United States |
| Elena DeSanti |
Brown University School of Public Health |
United States |
| Tom Alexander |
Purdue University Global |
United States |
Day 1: Virtual-Keynote Speakers:
| Denis Larrivee |
Loyola University Chicago |
United States |
| Elizabeth Dale Gilley |
The Elle Foundation |
United States |
Day 1: Virtual-Speakers:
| Ange Weinrabe |
The University of Sydney |
Australia |
| Santhosh Kumar J |
Amrita College of Nursing, Amrita Vishwa Vidyapeetham |
India |
| Sindu Padmanabhan |
Bharathiar University |
India |
| Moshe Bensimon |
Bar Ilan University |
Israel |
| Sushil Jha |
Jawaharlal Nehru University |
India |
| Paul Raj |
Jyoti Nivas College |
India |
| Ramesh Nagarajappa |
The Oxford Dental College at Bangalore |
India |
| Traci A Owens |
Attorney at Law |
United States |
| Jeffrey Lozon |
Bar Ilan University |
Israel |
| Michele M. Mahr |
California State University |
United States |
| Robert Paul Maddox II |
University of Wyoming |
United States |
| Juan Sanabri |
California State University |
United States |
| Hayrunnisa Unlu |
Mayo Clinic Arizona |
United States |
| Kelsey Whitus |
Drexel University College of Medicine |
United States |
| Thersilla Oberbarnscheidt |
University of Pittsburgh |
United States |
| Keith Klostermann |
State University of New York at Fredonia |
United States |
| Matthew Hanauer |
CleanSlate Centers |
United States |
| Matthew J Beattie |
University of Oklahoma |
United States |
| Joey Pagano |
Author, SPHS, The Traveling Social Worker |
United States |
| Cornel Stanciu |
New Hampshire Hospital |
United States |
| Brandon Lucke Wold |
University of Florida |
United States |
| Sara Haddadi |
University of Miami Miller |
United States |
| Baitubaev Dyussengali |
Psychiatrist-Narcologist |
Kazakhstan |
Day 1: Virtual-Posters:
| Arieh S Solomon |
Tel Aviv University |
Israel |
| Ghaith Adi |
Alfaisal University |
Saudi Arabia |
| Nikita Mehdiratta |
Smell and Taste Treatment & Research Foundation Ltd |
United States |
| Hyun Sue Kim |
Virginia Tech Carilion School of Medicine |
United States |
| Nikita Mehdiratta |
Smell and Taste Treatment & Research Foundation Ltd |
United States |
| Reshma Paul |
Cooper Medical School of Rowan University |
United States |
Day 2: In-Person-Keynote Speakers:
| Ann Marie Leonard Zabel |
Curry College & NEALAC Clinic |
United States |
| Carroy Ferguson |
University of Massachusetts-Boston |
United States |
| Roy F Baumeister |
Harvard University |
United States |
| Meera Vaswani |
All India Institute of Medical Sciences (AIIMS) |
India |
Day 2: In-Person- Speakers:
| Kimberley Berlin |
Compassionate Beginnings, LLC |
United States |
| Richard I. Suarez |
Florida International University |
United States |
| Martin Makela |
University of Washington |
United States |
| Sparkles Ransom |
Licensed Clinical Social Worker |
United States |
| Stephanie Cross |
WA Department of Children, Youth, and Families |
United States |
| Yihong Yang |
National Institute of Health |
United States |
| Alyssa Nieves |
Cal State University San Marcos |
United States |
| Izabela Saraiva |
Hospital de Clinicas de Porto Alegre |
Brazil |
| Ashton Christopher |
Center for Recovery |
Canada |
| Roshni Gandhi |
Cooper Medical School of Rowan University |
United States |
| Katherine Reavis |
University of South Florida |
United States |
| Catherine M Cahill |
Massachusetts General Hospital and Harvard Medical School |
United States |
| Jack Rogers |
Massachusetts General Hospital |
United States |
| Grace Hajinazarian |
Tufts University, School of Medicine |
United States |
| Berhanie Getnet Gebresilus |
University of Gondar |
Ethiopia |
| Raj Gopalan |
BSRM Consulting |
United States |
Day 2: Virtual-Keynote Speakers:
| Ken Ware |
NeuroPhysics Therapy Institute and Research Centre |
Australia |
| Dawn Duhaime |
Spring Green Foundation |
United States |
| Jun Hua |
Johns Hopkins University School of Medicine |
United States |
| David Sperbeck |
Private Practice |
United States |
Day 2: Virtual-Speakers:
| Kam Wilkinson |
Ken Ware NeuroPhysics Therapy |
Australia |
| Zhifang Xu |
Tianjin University of Traditional Chinese Medicine |
China |
| Lawrence Best |
Oxford University Hospitals NHS Trust |
United Kingdom |
| Nikita Sharma |
Swami Vivekanand Subharti University |
India |
| Christina Bitsara |
University of Cambridge |
United Kingdom |
| Buket Yilmaz |
Nizip Public Hospital |
Turkey |
| Merve Turkmen |
Sanko University Medical Faculty Hospital |
Turkey |
| Stanislav Engel |
Ben-Gurion University |
Israel |
| Mohamed Fathi Al Gharyani |
Benghazi Medical Center |
Libyan Arab Jamahiriya |
| Rotem Saar Ashkenazy |
Ashkelon Academic College |
Israel |
| Anne Dorothee Rosch |
University of Edinburgh |
United Kingdom |
| Daniel Clayman |
University of Nottingham |
United Kingdom |
| Cristian Ravariu |
Universitatea Politehnica Bucuresti |
Romania |
| Hesham Elnazer |
University of Sussex |
United Kingdom |
| Radwa Awad |
University of Texas Health Science Center at Houston (UTHealth) |
United States |
| David Zeng |
Johns Hopkins University |
United States |
| Marina Martinez-Vargas |
Baylor College of Medicine |
United States |
| Isabella Kim |
Academy of the Holy Angels |
United States |
| Kimberley D Ryan |
Brandon University |
Canada |
| Brandon Lucke Wold |
University of Florida |
United States |
| Aya El-Taibany |
Wake Forest Institute of Regenerative Medicine |
United States |
Day 3: In-Person-Keynote Speakers:
| Juan Moreira |
CNC/Gnosis Neurointegrative Center |
Puerto Rico |
| Yong Xiao Wang |
Albany Medical College |
United States |
| Jingsen Ma |
Dynaflow, Inc |
United States |
Day 3: In-Person-Speakers:
| Maryam Shahab |
Central Park Physicians |
United States |
| Nanxia Zhao |
Biogen |
United States |
| Ilse Saldivar |
Pain and Headache Centers of Texas |
United States |
| Ariyaneh Nikbin |
Albert Einstein College of Medicine |
United States |
| Ifechukwude J. Biose |
Louisiana State University Health Sciences Center |
United States |
| Alex Goraltchouk |
Remedium Bio |
United States |
| Ram Prajit |
Khon Kaen University |
Thailand |
| Fernanda Cristina Poscai Ribeiro |
Universidade do Oeste Paulista |
Brazil |
| Muhammad Yahya Saif |
Aston University |
United Kingdom |
| Nikita Dawan |
King Georges's Medical University |
India |
| Pallavi Chatterjee |
Saha Institute of Nuclear Physics |
India |
| Christine Akumcha Tekum |
Maarif International Schools of Equatorial Guinea |
Equatorial Guinea |
| Andres Villegas-Lanau |
Universidad de Antioquia |
Colombia |
| Faii Ong |
GyroGear |
United Kingdom |
Day 3: Virtual-Keynote Speakers:
| W S El Masri |
Keele University |
United Kingdom |
| Sam Vaknin |
Southern Federal University |
Russian Federation |
| Rocco J Gennaro |
University of Southern Indiana |
United States |
Day 3: Virtual-Speakers:
| Yang Du |
Shanghai Jiao Tong University School of Medicine |
China |
| Temitope Labinjo |
University of Sheffield |
United Kingdom |
| Vijayan Gurumurthy Iyer |
Bihar Institute of Public Administration & Rural Development |
India |
| Soewadi |
Gadjah Mada University |
Indonesia |
| Arda Ozkurt |
Hisar School |
Turkey |
| Georgios Matis |
University of Cologne |
Germany |
| Lama Saad El-Din Mahmoud |
October 6 University |
Egypt |
| Nico Morales |
No Halo LLC |
United States |
| Alphonsus Obayuwana |
Triple-H Project LLC |
United States |
| Vincent Colucciello |
Greenville School of Medicine |
United States |
| Sagarika Gopalkrishnan |
Einstein Medical Centre |
United States |
| Radika Saigal |
The Royal Wolverhampton Trust |
United Kingdom |
| Farsana Farooq |
Digital University Kerala |
India |
| Anita V. Handore |
Phytoelixir Pvt.Ltd |
India |
Day 3: Virtual-Posters:
| Jinwon Chang |
Korean Minjok Leadership Academy |
Korea |
| Aaron Kim |
Seoul International School |
Korea, Republic of |
| Anacleto Clent Banaay |
Southwestern University PHINMA School of Medicine |
Philippines |
| Huicong Wang |
Xuanwu Hospital of Capital Medical University |
China |
| Erika Jasukaitiene |
Lithuanian University of Health Sciences |
Lithuania |
| Drake Shafer |
California University of Science and Medicine |
United States |
| Seham Azzam |
TTUHSC School of Medicine |
United States |
| Badriah Alayidi |
University of Nottingham |
United Kingdom |
| Alicia Wells |
University of California, Irvine School of Medicine |
United States |
| Jaskeerat Gujral |
University of Pennsylvania |
United States |
| Stephen Avila |
Cedars Sinai Medical Center |
United States |
| Yaman Dalati |
Michigan State University |
United States |
| Aliaa Mousa |
Capital Health |
United States |
| Jane Zebrack |
Duke University |
United States |
| Priya Joshi |
Rowan-Virtua SOM |
United States |
| Johny Tran |
Temecula Valley Hospital |
United States |
| Saniya Ahmed |
West Virgnia School of Osteopathic Medicine |
United States |
| Swetha Ravi |
Michigan State University College of Osteopathic Medicine |
United States |
Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference | Brain Congress 2024 | International Neuroscience Conference 2024
We once again thank all the participants for their wonderful involvement towards the event which helped us for successful execution of this event.
Upcoming Neurology Conferences:
Conference Name: 9th Edition of International Conference on Neurology and Neurological Disorders (Hybrid Event)
Dates: June 20-22, 2024
Venue: Paris, France
Conference Name: 10th Edition of International Conference on Neurology and Brain Disorders (Hybrid Event)
Dates: October 21-23, 2024
Venue: Baltimore, Maryland, USA
Upcoming Addiction Conference:
Conference Name: 5th Edition of Global Conference on Addiction Medicine, Behavioral Health and Psychiatry (Hybrid Event)
Dates: October 21-23, 2024
Venue: Baltimore, Maryland, USA
For Magnus Group Conferences Reviews:
Magnus Group Neurology Conferences Reviews | Magnus Conferences Reviews
Details of Upcoming Neurology Conferences:
6th Edition of International Conference on Neurology and Neurological Disorders 2022 Report:
Magnus Group joyfully announces the successful conclusion of the "6th Edition of International Conference on Neurology and Neurological Disorders" (INBC 2022) held in Orlando, USA from October 24-26, 2022.
The congress revolved around the theme "Exploring the Grey Areas of Neurological Research."
INBC 2022 served as an advanced platform for the presentation and discussion of the latest discoveries and clinical practices in brain care. It addressed strategies for diagnosis, monitoring, and management to enhance neurodevelopmental outcomes. The conference delved into methods to understand, monitor, protect, and provide care for both developing and developed brains. Over the course of three days, the event offered valuable opportunities for continuing medical education to a diverse array of professionals dedicated to improving neurological and developmental health outcomes for neuro patients.
For INBC 2022 Final Program: Click Here
For INBC 2022 Abstract Book: Click Here
For INBC 2022 Gallery: Click Here
We express our heartfelt appreciation to the Keynote Speakers, Speakers, Moderators, Poster Presenters, Students, and Delegates for their active participation and valuable contributions, which significantly contributed to the success of the event.
We trust that you found satisfaction in both the scientific discussions and networking opportunities provided during the event. We hope that this experience has been beneficial in advancing your professional endeavors. We look forward with optimism to the prospect of continued collaboration with the majority of you in the near future.
Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
Organizing Committee INBC 2022:
| Ken Ware |
NeuroPhysics Therapy Institute and Research Centre |
Australia |
| Roy G Beran |
School of Medicine at Griffith University |
Australia |
| Thomas J Webster |
Interstellar Therapeutics |
United States |
| W S El Masri |
Keele University |
United Kingdom |
| Andreas Till |
University of Bonn |
Germany |
| Harinder Jaseja |
NIMS University |
India |
Magnus Group is prerogative to thank the Organizing Committee Members, Keynote Speakers; Chair and Co-chairs on engross the plenary sessions, workshops, and special sessions in an expanded manner to make this conference a privileged Summit.
INBC 2022 Speaker Line Up:
Day 1: Keynote Speakers
| Thomas J. Webster |
Interstellar Therapeutics |
United States |
| Ken Ware |
NeuroPhysics Therapy Institute |
Australia |
| Andreas Till |
University of Bonn |
Germany |
| W S El Masri |
Keele University |
United Kingdom |
Day 1: Speakers
| Fulya Turker |
Johns Hopkins University |
United States |
| Arman Fijany |
TCU School of Medicine |
United States |
| Marina Martinez-Vargas |
Baylor College of Medicine |
United States |
| Arman Fijany |
TCU School of Medicine |
United States |
| Amir Hadanny and Mohammed Elamir |
Aviv Clinics |
United States |
| Ulrich Sprick |
Alexius/Josef Clinic Neuss |
Germany |
| Aygun BadaLova |
University College London |
United Kingdom |
| Dell G Mars |
Southeastern Louisiana University |
United States |
| Lakshmi Sundeep |
Hindustan Institute of Technology and Science |
India |
| Xiaodong Cheng |
Shanghai University of Traditional Chinese Medicine |
China |
Day 1: Posters
| Alsu Zagorulko |
Illinois Center for Neurological and Behavioral Medicine |
United States |
| Solomon Nittala |
Larkin Community Hospital Palm Springs Campus |
United States |
| Elita Delbruck |
California University of Science and Medicine |
United States |
| Luis A. Sierra |
Beth Israel Deaconess Medical Center |
United States |
| Rebecca Lees |
Guthrie Robert Packer Hospital |
United States |
| Siri Tummala |
TCU School of Medicine |
United States |
| Gerald W. Grass |
Ketamine Research Institute |
United States |
| Maral Kasiri |
University of California |
United States |
| Johanna Christina Reiners |
Ulm University |
Germany |
| Judith Stefanie Scheller |
Ulm University |
Germany |
| Smaili Rachid |
University hospital of Tangier-Tétouan-Al Hoceima |
France |
| Peter Facchini |
Enveric Biosciences |
Canada |
| Jillian Hagel |
Enveric Biosciences |
Canada |
| Ilhan Yu |
Nowon eulji university hospital |
Korea, Republic of |
| Min-sung Gee |
Kyunghee University |
Korea, Republic of |
| Vildan Tuncbilek |
Tekirdag Namik Kemal University |
Turkey |
Day 2: Keynote Speakers
| Roy G Beran |
Western Sydney University |
Australia |
Day 2: Speakers
| Georgios Matis |
University Cologne Hospital |
Germany |
| Brandon Lucke-Wold |
University of Florida |
United States |
| Leya Maliekal |
Wayne State University School of Medicine |
United States |
| Flavia I. Spiroiu |
McMaster University |
Canada |
| David Chang |
NHS Hull University Teaching Trust |
United Kingdom |
| Alice Tebboth |
East Sussex Healthcare NHS Trust |
United Kingdom |
| Cristian Ravariu |
Polytechnic University of Bucharest |
Romania |
| Harinder Jaseja |
NIMS University |
India |
| Kunal Bhanot and Muhesh Taheem |
Imperial College London |
United Kingdom |
| Paul Raj |
Jyoti Nivas College |
India |
| Maria Joao Sacadura |
Catholic University of Portugal |
Portugal |
| Manuel Narvaez Pelaez |
University of Malaga |
Spain |
| Irene Fasciani |
University of L'Aquila |
Italy |
| Julia Souza E Costa |
Anhembi Morumbi University |
Brazil |
| Ana Beatriz Lima Pedroza |
Universidade Anhembi Morumbi |
Brazil |
| Noor Azzizah Omar |
National University of Malaysia |
Malaysia |
| Nishi M Satish |
VMMC and Safdarjung Hospital |
India |
| Keith Stenning |
University of Edinburgh |
Scotland |
Seyyedeh Somayyeh
Moshir Estekhareh |
Ferdowsi University of Mashhad |
Iran |
Day 2: Posters
| Ryane Elizabeth Adams |
McGovern Medical School |
United States |
| Jacob Saucier |
University of Sherbrooke |
Canada |
| Beata Lindholm |
Lund University |
Sweden |
| Gabriela Dumitrita Stanciu |
Grigore T. Popa University of Medicine and Pharmacy |
Romania |
| Mariam Al-Umran |
Alfaisal University |
Saudi Arabia |
| Begum Bulgurluoglu |
Hisar Schools |
Turkey |
| Marcia Castillo |
Centro de Salud Fundación Activo 20-30 |
Dominican Republic |
| Zhi-Hao Liu |
Daping Hospital |
China |
| Khadga Raj Aran |
ISF College of Pharmacy |
India |
| Vanessa Veronica |
Duta Wacana Christian University |
Indonesia |
Day 3: Keynote Speakers
| Torbjorn Backstrom |
University of Umea |
Sweden |
Day 3: Speakers
| Mohammad Abu-Abaa |
Capital Health Regional Medical Center |
United States |
| Geetanjali Rathore |
University of Nebraska Medical Center |
United States |
| Benjamin Pinker |
Imperial College London |
United Kingdom |
| Ana Lilia Rodríguez Villegas |
Universidad Nacional Autónoma de México |
Mexico |
| Tongtong Li |
Michigan State University |
United States |
| Yasemin Tugce Yayla |
Memorial Sisli Hospital |
Turkey |
| Michelle Herman |
Michelle Herman Speaks |
United States |
| Annapurna Ahuja |
Hazaribag College of Dental Sciences and Hospital |
India |
| Almutazballlah Bassam Qablan |
Jordan University of Science and Technology |
Jordan |
| Paulo Henrique Leite Souza |
Nove de Julho University |
Brazil |
| Priyanka Sethi |
Manav Rachna International Institute of Research and Studies |
India |
| Bruany Antoniolli Bianchi |
Universidade Anhembi Morumbi |
Brazil |
| Lama Saad El-Din Mahmoud |
October 6 University |
Egypt |
| Esraa Askar |
Northwell/Hofstra Zucker School of Medicine |
United States |
| Fareha Khalil |
University of Limerick |
Ireland |
| Hamza Islam |
Punjab Medical College |
Pakistan |
| Rabia Islam |
Punjab Medical College |
Pakistan |
| Sajjad Saghebdoust |
Tehran University of Medical Sciences |
Iran |
Day 3: Posters
| Shreenal Patel |
HCA healthcare Houston |
United States |
| Hannah Girgis |
Aultman Hospital |
United States |
| Joshua Burshtein |
Donald and Barbara Zucker School of Medicine |
United States |
| David Galel |
Rosalind Franklin University of Medicine and Science |
United States |
| Neil Gerts |
Los Robles Regional Medical Center |
United States |
| Rebecca Simon |
Nova Southeastern University |
United States |
| Mohammad Abu-Abaa |
Capital Health Regional Medical Center |
United States |
| Patrick Brown |
The University of Queensland |
United States |
| Anthia Papazoglou |
Cantonal Hospital Aarau |
Switzerland |
| Pourya Bahiraei |
Tehran University of Medical Sciences |
Iran |
Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
We once again thank all the participants for their wonderful involvement towards the event which helped us for successful execution of this event.
For Magnus Group Conferences Reviews:
Magnus Group Neurology Conferences Reviews | Magnus Conferences Reviews
4th Edition of International Conference on Neurology and Neurological Disorders 2021 Report:
Magnus Group would like to thank everyone who attended “4th Edition of International Conference on Neurology and Neurological Disorders” which is going to be held Virtually during September 09 -10, 2021. The Paricipants from different nations volunteered their time and resources to make the event a success.
We would like to express our sincere thanks to Keynote Speakers, Speakers, Moderators, Poster Presenters, Students and delegates for attending and making the event a successful one.
Hopefully, you enjoyed both the scientific and networking aspects of the event and took use of the opportunity to expand your existing profession. I am optimistic that the vast majority of you will continue to work with me in the near future.
Check our website for regular updates: https://neurologycongress.com/
For INBC 2021 Final Program: Click Here
For INBC 2021 Abstract Book: Click Here
The conference aimed with a theme “Advancements and challenges in Neuroscience & Brain Disorders”. The summit engrossed a vicinity of sensible discussions on subjects like Neurodegenerative disorders, Behavioural Neurology, Neuropsychiatry, Paediatric Neurology, Neurological Disorders and Stroke, Neuroimmunology and Neurological Infections, Epilepsy & Seizure Disorders. The three days event implanted a firm relation of upcoming strategies in the field of Neurology, Neurodegenerative Disorders and Neurological Disorders and Stroke with the scientific community. The conceptual and valid knowledge shared, will also raise organizational collaborations to develop scientific accelerations.
Organizing Committee INBC 2021:
| Ken Ware |
Neuro Physics Therapy Institute and Research Centre |
Gold Coast, Queensland, Australia |
| Svyatoslav Medvedev |
Institute of the Human Brain RAS |
Russian Federation |
| Alexies Dagnino |
Universidad de Valparaíso |
Valparaíso, Chile |
Magnus Group is prerogative to thank the Organizing Committee Members, Keynote Speakers; Chair and Co-chairs on engross the plenary sessions, workshops, and special sessions in an expanded manner to make this conference a privileged Summit.
The conference was boarded with an opening ceremony followed by a series of lectures delivered by both Honorable Guests and members of the Keynote forum. The best Part of the conference were the keynote forum by prominent scientists, Ken Ware, Neuro Physics Therapy Institute and Research Centre, Gold Coast, Queensland, Australia; Svyatoslav Medvedev, Institute of the Human Brain RAS, Russian Federation; Alexies Dagnino, Universidad de Valparaíso, Valparaíso, Chile; gave their profitable contributions in the form of highly enlightening presentations and made the conference a best notch one.
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
INBC 2021 Speaker Line Up:
Day 1: Speakers
| Raffaele Pilla |
John of God Hospital – Fatebenefratelli |
Benevento, Italy |
| Vanessa Douet-Vannucci |
University of Côte-d’Azur University |
France |
| Wen-Chang Li |
School of Psychology and Neuroscience |
University of St Andrews, Uk |
| Gaurav Jha |
Queen’s Hospital |
BHRUT, UK |
| Wilfried Schupp |
Friedrich-Alexander University Erlangen-Nuremberg |
Germany |
| Vir B Singh |
Albany College of Pharmacy and Health Sciences |
USA |
| M Dubois Jeremy |
National Institute of Health and Medical Research |
France |
| Lama Saad El-Din Mahmoud |
October 6 University |
Egypt |
| Hussein Akel |
Institute of Pharmaceutical Technology and Regulatory Affairs |
Hungary |
| Alba Ávila Suárez |
Universidad del Desarrollo |
Chile |
Day 1: Posters
| Maryam Farhang |
Millennium Institute for Depression and Personality Research |
Chile |
| Marta Czajkowska |
University of Warmia and Mazury |
Poland |
| Polina Sobolevskaia |
Saint Petersburg State University |
Russia |
| Yahia Al Turk |
Henry Ford Hospital |
USA |
| Ali Hammed |
Tishreen University Hospital |
Syria |
| Pacôme Kouadio N’Go |
Pelefero Gon Coulibaly University |
Ivory Coast, Morocco |
| Ayushi Shah |
Kemper House |
USA |
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
Day 2: Speakers
| Sven-Erik Eriksson |
Division of Neurology |
Sweden |
| Alberto A. Rasia-Filho |
Federal University of Health Sciences of Porto Alegre |
Brazil |
| Georgina Rodríguez de Lores Arnaiz |
Institute of Cell Biology and Neurosciences |
Argentina |
| Silvia Lores Arnaiz |
University of Buenos Aires |
Argentina |
| Glòria Garrabou |
IDIBAPS - CIBERER - UB |
Spain |
| Meenal Dhall |
University of Delhi |
India |
| Balakrishnan Arivalagan |
Command Hospital Air Force |
India |
| Ali Imran |
Government College University |
Pakistan |
| Silvana Giuliatti |
University of Sao Paulo |
Brazil |
| Hussein Jaafar Hussein Ahmed |
University of Khartoum |
Sudan |
Day 2: Posters
| Betsy Dang |
Liverpool Hospital |
Australia |
| Bilal Vanlioglu |
Liverpool Hospital |
Australia |
| Maria V Soloveva |
Monash University |
Australia |
| Anna Litwiniuk |
Centre of Postgraduate Medical Education |
Poland |
| Zuzana Capkov |
University Hospital Olomouc |
Czech Republic |
| Julia Boytsova |
Bechtereva Institute of the Human Brain |
Russia |
| Maria Joao Cabral Sacadura |
Catholic University of Portugal and University of Coimbra |
Portugal |
| Nipuna Thivanka Weerasinghe |
National Hospital Kandy |
Srilanka |
| Mohamad Ayajuddin |
Nagaland University |
India |
We once again thank all the participants for their wonderful involvement towards the event which helped us for successful execution of this event.
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
For Magnus Group Conferences Reviews:
Magnus Group Neurology Conferences Reviews | Magnus Conferences Reviews
International Conference on Neurology and Brain Disorders 2019 Report:
Magnus Group takes great pride in announcing the success of “3rd Edition of International Conference on Neurology and Brain Disorders” (INBC 2019) which was held in Paris, France during June 24-26, 2019.
Neurology Congress 2019 witnessed a combination of peerless speakers who enlightened the crowd with their knowledge and confabulated on various new fangled issues related to the field of Neurology. The extremely illustrious conference hosted by Magnus Group was marked with the attendance of young and brilliant researchers, business delegates and talented student communities representing their countries around the world.
For INBC 2019 Final Program: Click Here
For INBC 2019 Abstract Book: Click Here
For INBC 2019 Gallery: Click Here
Organizing Committee INBC 2019:
| Aage R Moller |
The University of Texas at Dallas School of Behavioral and Brain Sciences |
USA |
| Ken Ware |
NeuroPhysics Therapy Institute |
Australia |
| Sam Vaknin |
Southern Federal University |
Israel |
| Marat Akhmet |
Middle East Technical University |
Turkey |
| Margarit Tadevosyan |
Artmed Medical Rehabilitation Center |
Armenia |
| Reidun Stenberg |
University Hospital Research Center |
Sweden |
| Zdzislaw Chilmonczyk |
National Medicine Institute |
Poland |
The Organizing Committee would like to thank the moderators Pooja George, Hamad General Hospital, Qatar and Susan Hawthorne, Ulster University, United Kingdom for their offerings which resulted in smooth functioning of the conference.
The conference was boarded with an opening ceremony followed by a series of lectures delivered by both Honorable Guests and members of the Keynote forum. The best Part of the conference were the keynote forum by prominent scientists, Ken Ware, NeuroPhysics Therapy Institute and Research Centre, Australia; Marat Akhmet, Middle East Technical University, Turkey; Stephen Grossberg, Boston University, USA; Reidun Stenberg, University Hospital Research Center, Sweden; Margarit Tadevosyan, “Artmed” Medical Rehabilitation Center, Armenia; William S Baek, Parkside Medical Group, USA gave their profitable contributions in the form of highly enlightening presentations and made the conference a best notch one.
Magnus Group is prerogative to thank the Organizing Committee Members, Keynote speakers, Chair and Co-chairs on engross the plenary sessions, workshops, and special sessions in an expanded manner to make this conference a privileged Summit.
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
INBC 2019 Speaker Line Up:
Day 1: Speakers
| Mary Payne |
Marshall University School of Medicine |
USA |
| Natasa Dragicevic |
University of Connecticut Health Center |
USA |
| Jeffrey Edwards |
Brigham Young University |
USA |
| Mojgan Rastegar |
University of Manitoba |
Canada |
| Valida Isanova |
Kazan State Medical University |
Russian Federation |
| Richard Rokyta |
Charles University in Prague |
Czech Republic |
| Yan Wang |
Chengdu Fifth People’s Hospital Chengdu |
China |
| Csokay Andras |
Military Hospital |
Hungary |
| Umur Kayabasi |
Bahcesehir University |
Turkey |
Day 2: Speakers
| Ken Ware |
NeuroPhysics Therapy Institute and Research Centre |
Australia |
| Aage R moller |
The University of Texas at Dallas School of Behavioral and Brain Sciences |
USA |
| Christian C. Naus |
University of British Columbia |
Canada |
| Andre Eduardo de Almeida Franzoi |
University of the Region of Joinville |
Brazil |
| Anna Konopka |
Macquarie University |
Australia |
| Joe Herbert |
University of Cambridge |
UK |
| Janet Leathem |
Massey University |
New Zealand |
| Husham Bayazed |
University of Zakho |
Iraq |
| Valentina Barkhanskaya |
Karaganda Medical University |
Kazakhstan |
Day 2: Poster Presenters:
| Susan Hawthorne |
Ulster University |
United Kingdom |
| Yida Wang |
Georgetown University Medical Center and Washington Institute for Health Sciences |
USA |
| Nilgun Cinar |
Maltepe University Hospital |
Turkey |
| Kun Xiong |
Xiangya Medical School Central South University |
China |
| Hee-Young Kim |
Pusan National University |
Republic of Korea |
| Ji-yoon Lee |
Daejeon University |
Republic of Korea |
| Ping Heng Tan |
Chi Mei Medical Center |
Taiwan |
| Hea Eun Yang |
Veterans Health Service Medical Center |
Republic of Korea |
| Pooja George |
Hamad General Hospital |
Qatar |
| Hamid Sami |
Quinnipiac University |
USA |
| Fernanda Bonilla Colome |
Faculdades Pequeno Principe |
Brazil |
| Moncy Thomas |
Cleveland Clinic Abu Dhabi |
UAE |
| Fernanda Bonilla Colome |
Faculdades Pequeno Principe |
Brazil |
Day 3: Speakers
| Ramel Carlos |
The Neurology Clinic |
USA |
| Jinfeng Yang |
Central South University |
China |
| Boulenour Mesraoua |
Weill Cornell Medical College |
Qatar |
| Neil Jeyasingam |
Sydney University |
Australia |
| Bengt Sivberg |
Lund University |
Sweden |
| Elnura Eralieva |
Osh City Clinical Hospital |
Kyrgyzstan |
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
For Magnus Group Conferences Reviews:
Magnus Group Neurology Conferences Reviews | Magnus Conferences Reviews
2nd Edition of International Conference on Neurology and Brain Disorders 2018 Report:
Magnus Group takes a great pride in announcing the “2nd Edition of International Conference on Neurology and Brain Disorders (INBC 2018)" which was held in Rome, Italy during June 04-06, 2018.
For INBC 2018 Final Program: Click Here
For INBC 2018 Abstract Book: Click Here
For INBC 2018 Gallery: Click Here
The conference aimed with a theme “Advancements and Challenges in Neurosciences & Brain Disorders”. The summit engrossed a vicinity of sensible discussions on subjects like Neurodegenerative disorders, Behavioural Neurology, Neuropsychiatry, Paediatric Neurology, Neurological Disorders and Stroke, Neuroimmunology and Neurological Infections, Epilepsy & Seizure Disorders. The three days event implanted a firm relation of upcoming strategies in the field of Neurology, Neurodegenerative Disorders and Neurological Disorders and Stroke with the scientific community. The conceptual and valid knowledge shared, will also raise organizational collaborations to develop scientific accelerations.
Organizing Committee INBC 2018:
| Stephen Grossberg |
Boston University |
USA |
| Harry W M Steinbusch |
Maastricht University |
Netherlands |
| P Hemachandra Reddy |
Texas Tech University Health Sciences |
USA |
| Anthony E. Kline |
University of Pittsburgh |
USA |
| Corina O. Bondi |
University of Pittsburgh |
USA |
| Maria-Magdalena Georgescu |
Louisiana State University |
USA |
| Marat Akhmet |
Middle East Technical University |
Turkey |
| Ken Ware |
NeuroPhysics Therapy Institute and Research Centre |
Australia |
| Ichiro Maruyama |
Okinawa Institute of Science and Technology Graduate University |
Japan |
| Fuyong Jiao |
Xian Jiaotong Univeristy |
China |
| Margarit Tadevosyan |
Artmed Medical Rehabilitation Center |
Armenia |
| Medvedev Svyatoslav |
N.P.Bechtereva Institute of the Human Brain of the Russian academy of sciences |
Russisa |
| Najib Murr |
Southern Illinois University School of Medicine |
USA |
The conference was boarded with an opening ceremony followed by a series of lectures delivered by both Honorable Guests and members of the Keynote forum. The best Part of the conference was the keynote forum by prominent scientists, Maria-Magdalena Georgescu, Louisiana State University, USA; Stephen Grossberg, Boston University, USA; Ken Ware, NeuroPhysics Therapy Institute and Research Centre, Australia; Corina O. Bondi, University of Pittsburgh, USA; Ichiro Maruyama, Okinawa Institute of Science and Technology Graduate University, Japan; Margarit Tadevosyan, Artmed Medical Rehabilitation Center, Armania; Svyatoslav Medvedev, N.P. Bechtereva Institute of the Human Brain of the Russian Academy of Sciences, Russia; P. Hemachandra Reddy, Texas Tech University Health Sciences, USA; Corina O. Bondi, University of Pittsburgh, USA; Najib Murr, SIU Medicine, USA; Viviane Elsner, IPA Methodist University, Porto Alegre-RS Brazil; Marat Akhmet, Middle East Technical University, Turkey; gave their profitable contributions in the form of highly enlightening presentations and made the conference a best notch one.The Organizing Committee would like to thank the moderators Jodie Marquez, University of Newcastle, Hunter Medical Research Institute, Australia, Lucia Carvelli, Florida Atlantic University, USA for their contributions which ensued in smooth functioning of the conference.
Magnus Group is prerogative to thank the Organizing Committee Members, Keynote speakers, Chair and Co-chairs on engross the plenary sessions, workshops and special sessions in a expanded manner to make this conference an privileged Summit.
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
INBC 2018 Speaker Line Up:
Day 1 Speakers:
| Stephen Grossberg |
Boston University |
USA |
| Ken Ware |
NeuroPhysics Therapy Institute and Research Centre |
Australia |
| Maria-Magdalena Georgescu |
Louisiana State University |
USA |
| Corina O. Bondi |
University of Pittsburgh |
USA |
| Harry W.M. Steinbusch |
Maastricht University Medical Center |
Netherlands |
| Floyd Thompson |
Brain Rehabilation Center |
USA |
| Luca Malfassi |
La Cittadina Fondazione Studi e Ricerche Veterinarie |
Italy |
| Karol S. Bruzik |
The University of Illinois at Chicago |
USA |
| Swetlana Sirko |
Ludwig-Maximilians-University Munich |
Germany |
| Vittorio Maglione |
IRCCS Neuromed |
Italy |
| Valida Isanova |
Kazan State Medical University |
Russia |
| Jodie Marquez |
University of Newcastle |
Australia |
| Anthony E. Kline |
University of Pittsburgh |
USA |
Day 2 Speakers:
| P. Hemachandra Reddy |
Texas Tech University Health Sciences |
USA |
| Ichiro Maruyama |
Okinawa Institute of Science and Technology Graduate University |
Japan |
| Svyatoslav Medvedev |
N.P.Bechtereva Institute of the Human Brain of the russian academy of sciences |
Russia |
| Margarit Tadevosyan |
Artmed Medical Rehabilitation Center |
Armania |
| CarolAnn Peterson |
University of Southern California |
USA |
| Nicola Postol |
University of Newcastle |
Australia |
| Eleonora Palmieri |
University in Macerata |
Italy |
| Yasutoshi Koga |
Kurume University Graduate School of Medicine |
Japan |
| Eugenia V Gurevich |
Vanderbilt University |
USA |
| Arun L. Jayaraman |
Mayo Clinic Arizona |
USA |
Day 2 Posters:
| Richard Francis |
Royal united hospital |
UK |
| Baranowska-Bosiacka Irena |
Pomeranian Medical University in Szczecin |
Poland |
| Marta Goschorsk |
Pomeranian Medical University in Szczecin |
Poland |
| Izabela Gutowska |
Pomeranian Medical University in Szczecin |
Poland |
| Hiroko IKESHIMA-KATAOKA |
WASEDA University |
Japan |
| Hyun Im Moon |
Bundang Jesaeng Hospital |
Korea |
| Kruchinina Margarita Vitalievna |
Siberian Branch of the Russian Academy of Sciences |
Russia |
| Wasik Agnieszka |
Institute of Pharmacology PAS |
Poland |
| Cesar Garcia-Cruz |
Centro de Investigación y de Estudios Avanzados del IPN |
México |
| Carlos Eduardo Monici Fernandes |
Federal University |
Brazil |
| Bruna Silva Marques |
Federal University of Sao Carlos |
Brazil |
| Zofia Rogoz |
Institute of Pharmacology Polish Academy of Sciences |
Poland |
| Kelly Regina Serafim |
Center Universitary Central Paulista |
Brazil |
| Bartlomiej Stanczykiewicz |
Wroclaw Medical University |
Poland |
| Choon-Gon Jang |
Sungkyunkwan University |
Korea |
| Howard. H. Fenn |
Stanford University |
USA |
| Keerthi Madhurya Kethireddi |
East Sussex Healthcare NHS trust |
UK |
| Bernadete Marcia Voichcoski |
Technological Federal University of Parana/Electronica |
Brazil |
| Dao Pan |
Cincinnati Children’s Hospital Medical Center |
USA |
| Anastassiya Perfilyeva |
Institute of General Genetics and Cytology CS MES RK |
Kazakhstan |
| Leyla Djansugurova |
Institute of General Genetics and Cytology CS MES RK |
Kazakhstan |
| Karine Bertoldi |
Universidade Federal do Rio Grande do Sul |
Brazil |
| Arjay Dannug |
Makati Medical Center |
Phillipnes |
Day 3 Speakers:
| Najib Murr |
SIU Medecine |
USA |
| Viviane Elsner |
IPA Methodist University |
Porto Alegre-RS Brazil |
| Marat Akhmet |
Middle East Technical University |
Turkey |
| Lucia Carvelli |
Florida Atlantic University |
USA |
| Yu Liu |
Jinan University |
China |
| Maurizio Petrarca |
“Bambino Gesù” Children’s Hospital |
Italy |
| Patrice Ntenga |
Neurological Clinic of the National Teaching Hospital |
FANN |
| Lorena Rubio |
Universidad Nacional Autonoma de Mexico |
Mexico |
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
Delegates:
| Kimberley Naylor |
Central Otago Health Services Ltd |
New Zealand |
| Christian Saleh |
Rehab Basel Neuro |
Switzerland |
| INCHUL JUNG |
Daejeon University |
Korea |
| Ho Choong |
Campbell town Hospital |
Australia |
| Dina Dimitrova Bayatova |
Actavis EAD A Teva company |
Bulgaria |
| Galini Georgiev Hristov |
Actavis EAD A Teva company |
Bulgaria |
| RADIANA KOLEVA KOLEVA |
Actavis EAD A Teva company |
Bulgaria |
| Nadka Ivanova Samardzhieva |
Actavis EAD A Teva company |
Bulgaria |
| Svetlana Alexandrova Beneva |
Actavis EAD A Teva company |
Bulgaria |
| Hristina Spiridonova Chilingirova – Ignatieva |
Actavis EAD A Teva company |
Bulgaria |
| David |
CMDHB |
New Zealand |
| Andrew Tidemann |
Women's and Children's Hospital Adelaide |
Australia |
| MD MOSARRAF HOSSAIN |
Uttara Adhunik Medical College |
Bangladesh |
| Navid Tabibzadeh |
Lehigh Valley Health Network |
USA |
| JG Storosum |
AMC |
Netherlands |
| J Hoogendijk |
UMCU |
Netherlands |
| Heidi Muller |
TNU Aarhus University |
Denmark |
| Betina Elfving |
TNU Aarhus University |
Denmark |
| Armin Nikpour |
University of Sydney |
Australia |
| Jean Dore |
University of Laval Canada |
Canada |
2nd Edition of International Conference on Neurology & Brain Disorders would not have been successful if it has not been supported by intercontinental, multi-professional steering committee.
We once again thank all the participants for their wonderful participation towards the event which helped us for effective accomplishment of this event.
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
For Magnus Group Conferences Reviews:
Magnus Group Neurology Conferences Reviews | Magnus Conferences Reviews
International Conference on Neurology and Brain Disorders 2017 Report:
Magnus Group is delighted to declare the success of the "International Conference on Neurology and Brain Disorders (INBC 2017)", held in Valencia, Spain from June 26-28, 2017.
The Neurology Congress 2017 featured an exceptional lineup of speakers who shared their profound knowledge and engaged in discussions on innovative issues within the field of Neurology. This distinguished conference, hosted by Magnus Group, was characterized by the presence of young and accomplished researchers, business delegates, and talented student communities from various countries across the globe.
For INBC 2017 Final Program: Click Here
For INBC 2017 Abstract Book: Click Here
For INBC 2017 Gallery: Click Here
The conference aimed with a theme “Advancements and Challenges in Neurosciences & Brain Disorders”. The summit engrossed a vicinity of sensible discussions on subjects like Neurodegenerative disorders, Behavioural Neurology, Neuropsychiatry, Paediatric Neurology, Neurological Disorders and Stroke, Neuroimmunology and Neurological Infections, Epilepsy & Seizure Disorders. The three days event implanted a firm relation of upcoming strategies in the field of Neurology, Neurodegenerative Disorders and Neurological Disorders and Stroke with the scientific community. The conceptual and valid knowledge shared, will also raise organizational collaborations to develop scientific accelerations.
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
Organizing Committee INBC 2017:
| Giuseppe Scalabrino |
University of Milan |
Italy |
| Harry W.M. Steinbusch |
Maastricht University |
The Netherlands |
| Pankaj Sharma |
University of London |
UK |
| Henry Bakunts |
International Medical Centre “STROKE” |
Republic of Armenia |
| Mira Rakacolli-Kapisyzi |
President of Albanian Society of Neurology |
Albania |
| Serhiy Forostyak |
Charles University |
Czech Republic |
Magnus Group is prerogative to thank the Organizing Committee Members, Keynote speakers, Chair and Co-chairs on engross the plenary sessions, workshops, and special sessions in an expanded manner to make this conference a privileged Summit.
The conference was boarded with an opening ceremony followed by a series of lectures delivered by both Honorable Guests and members of the Keynote forum. The best Part of the conference were the keynote forum by prominent scientists, Sergi Ferre, National Institute on Drug Abuse(NID, NIH), USA; Marisela Morales, National Institute on Drug Abuse(NID, NIH), USA; Giuseppe Scalabrino, University of Milan, Italy; Harry W.M. Steinbusch, Maastricht University, The Netherlands; Pankaj Sharma, University of London, UK; Mira Rakacolli-Kapisyzi, President of Albanian Society of Neurology, Albania; Henry Bakunts, International Medical Centre “STROKE”, Armenia; gave their profitable contributions in the form of highly enlightening presentations and made the conference a best notch one.The Organizing Committee would like to thank the moderators Dr. Leonardo Pignataro, Columbia University-City University of New york-CSI, USA, Dr. Udai Pandey, University of Pittsburgh Medical Center, USA, Michael Ugrumov, Institute of Development Biology RAS, Russia for their offerings which resulted in smooth functioning of the conference.
INBC 2017 Speaker Line Up:
Day 1: Speakers
| Michel Baudry |
Western University of Health Sciences |
USA |
| Miranda N. Reed |
Auburn University |
USA |
| Stephen Wren |
University of Oxford |
UK |
| KHIN MAUNG BO |
Northern Lincolnshire and Goole NHS Foundation Trust |
UK |
| Mahmoud Kiaei |
University of Arkansas for Medical Sciences |
United States |
| Michael Ugrumov |
Institute of Developmental Biology RAS |
Russian Federation |
| Kimiko Inoue |
Toneyama National Hospital |
Japan |
| Caroline Corbel |
Institut de Recherche Dupuy de Lome (IRDL) |
France |
| Jong Wook Chang |
Samsung Medical Center |
Korea |
| Nicole Hess |
University of New England |
Australia |
| Udai Pandey |
University of Pittsburgh Medical Center |
USA |
| Gabriele Saretzki |
Newcastle University |
UK |
| Shinji Ohara |
Matsumoto Medical Center |
Japan |
| Cristine Alves da Costa |
Institut de Pharmacologie Moléculaire et Cellulaire |
France |
| Leonardo Pignataro |
Columbia University-City University of New York-CSI |
USA |
| Sabine Cordes |
Lunenfeld-Tanenbaum Research Institute/Mt Sinai Hospital |
Canada |
| Jeffrey Liddell |
University of Melbourne |
Australia |
| Niall Finnerty |
Maynooth University |
Ireland |
| Debashis Mukhopadhyay |
Saha Institute of Nuclear Physics |
India |
| Gilles Guillemin |
Macquarie University |
Australia |
| Abigail Takyi |
University of Brighton |
UK |
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
Day 2: Speakers
| Medvedev Svyatoslav |
N.P.Bechtereva Institute of the Human Brain of the Russian Academy of Sciences |
Russian Federation |
| Martin L. Pall |
Washington State University |
USA |
| Sergio Chieffi |
Second University of Naples |
Italy |
| Laura Calza |
University of Bologna |
Italy |
| Andrzej Pilc |
Polish Academy of Sciences |
Poland |
| Razvana Stanciu |
Universite Libre de Bruxelles (ULB) |
Belgium |
| Kathryn Commons |
Children’s Hospital Boston-Harvard Medical School |
USA |
| Caroline Lucke |
Medical Campus University of Oldenburg |
Germany |
| Sabrina Wang |
National Yang-Ming University |
Taiwan |
| Marta Nieto |
Spanish National Research Council |
Spain |
| Viviane Rostirola Elsner |
IPA Methodist University |
Brazil |
| K.L. Leenders |
University of Groningen |
The Netherlands |
| Leah K. Light |
Brainchild Institute |
USA |
| Robyn Tolhurst |
Red Fern Communication |
Australia |
| Kenneth Gaines |
Vanderbilt university Medical Center |
USA |
| Bruno Gonzalez |
Inserm - U1245 Team NeoVasc |
France |
| Raquel Sofia Marques Neves |
Amana Healthcare Medical and Rehabilitation Hospital |
UAE |
| Marina Vladimirovna Zueva |
Moscow Helmholtz Research Institute of Eye Diseases |
Russia |
| Laehyun Kim |
Korea Institute of Science and Technology |
Korea |
| Prokopenko Semen |
Krasnoyarsk State Medical University |
Russian Federation |
| Luyang Tao |
Soochow University |
China |
| Anna Bezdeneznykh |
Krasnoyarsk State Medical University |
Russian Federation |
| M.R. Graham |
Llantarnam Health Care |
UK |
| Serhiy Forostyak |
Charles University |
Czech Republic |
| Gladstone C McDowell |
Integrated Pain Solutions |
USA |
| Saema Ansar |
Lund University |
Sweden |
| Barbara R. Cardoso |
University of Sao Paulo |
Brazil |
| Hassan Ravari |
Mashhad university of medical sciences |
Iran |
We once again thank all the participants for their wonderful involvement towards the event which helped us for successful execution of this event.
Day 3: Speakers
| Lars Hakan Thorell |
Linkoping University |
Sweden |
| Martin Egerth |
Lufthansa Aviation Training GmbH |
Germany |
| Munzberg Mathias |
BG Klinik Ludwigshafen |
Germany |
| Cecilia Montanez |
Centro de Investigacion y de Estudios Avanzados del IPN |
Mexico |
| Moataz Mohamed Talaat Mohamed Kamel El Semary |
Cairo university |
Egypt |
| Maria-Magdalena Georgescu |
Louisiana State University |
USA |
| Teruna J. Siahaan |
The University of Kansas |
USA |
| Toshiki Mizuno |
Kyoto Prefectural University of Medicine |
Japan |
| Katherine L Wisner |
Northwestern University |
USA |
| Meena Kumari |
Kansas State University |
USA |
| Magnus Gram |
Lund University |
Sweden |
| Joanna Czarzasta |
University of Warmia and Mazury |
Poland |
| Dennis J. Dlugos |
University of Pennsylvania School of Medicine |
USA |
| Lynda El-Hassar |
Yale School of Medicine |
USA |
| Hoi Ki Kate Lui |
Tseung Kwan O Hospital |
Hong Kong |
| Shuhei Yamaguchi |
Shimane University |
Japan |
| Victor Vvedensky |
Kurchatov Inststute |
Russian Federation |
| Michael Luedtke |
Johnson & Johnson |
USA |
| Fatimah Alqarni |
King Abdullah bin Abdulaziz University hospital |
Saudi Arabia |
| Bilgehan Atilgan ACAR |
Sakarya University Faculty of Medicine |
Turkey |
| Albekov Nurvadi |
Chechen State University |
Russia |
| Paul Chapple |
Queen Mary University of London |
UK |
| Turkan ACAR |
Sakarya University Faculty of Medicine |
Turkey |
Recommended Neuroscience Conference 2024 | Neurology Conferences | Brain Disorders Conference | Neurology Conference 2024 | Neuroscience Conferences | Neurology Conference | Brain Disorders Conferences | Neuroscience Conference| Brain Congress 2024 | International Neuroscience Conference 2024
For Magnus Group Conferences Reviews:
Magnus Group Neurology Conferences Reviews | Magnus Conferences Reviews